SPHN RDF Schema
Scope of the SPHN RDF Schema
The SPHN RDF Schema provides an interoperable framework for conveying information and storing health-related data from SPHN-related projects using Semantic Web technologies including RDF (see background section on RDF). The schema facilitates the integration of existing external resources within the SPHN Semantic Interoperability Framework. The SPHN RDF Schema is based on the SPHN Dataset (available at: https://sphn.ch/document/sphn-dataset/) and transforms its elements into a formal structure (see Figure 1).
This documentation provides an overview of the content of the SPHN RDF Schema that complies with version 2024.2 of the schema. A visual representation of the schema can be found at: https://biomedit.ch/rdf/sphn-schema/sphn.
Figure 1. Core elements defined in the SPHN dataset and their translation into RDF.
Concepts of the SPHN dataset are translated into classes (owl:Class)
and concepts’ compositions become either object properties (owl:ObjectProperty)
or data properties (owl:DataProperty) depending on the type of the value.
Table 1 summarizes these properties.
Property |
Concept composition |
Example |
|---|---|---|
Object |
Another concept |
A class |
Qualitative element |
A set of values valid for a concept provided in the SPHN dataset |
|
Data |
String |
|
Quantitative |
|
|
Temporal |
|
|
URI |
|
Value sets defined in the dataset are represented as individuals (owl:NamedIndividual) with their own namespace.
Technical specification of the schema
SPHN RDF namespace
The namespace of the SPHN RDF Schema may be divided into three parts:
The SPHN RDF Schema IRI: https://biomedit.ch/rdf/sphn-schema/sphn. The SPHN RDF Schema IRI remains fixed and must be defined in the SPHN RDF Schema. The SPHN RDF Schema IRI can be considered as the “base prefix” and will be used by both data providers (to annotate data) and data users (to query for the relevant classes/properties).
The SPHN versioned IRI: https://biomedit.ch/rdf/sphn-schema/sphn/2024/2. It is provided by the DCC at each published release of the SPHN RDF Schema and enables to distinguish different versions of the SPHN RDF Schema. The versioned IRI is used by the projects to refer to a specific version of the SPHN RDF Schema and is therefore included in the header of all datasets generated using this schema version.
The SPHN RDF Schema individuals’ IRI: https://biomedit.ch/rdf/sphn-schema/sphn/individual#
Note
The SPHN RDF Schema has dereferenceable links meaning that the content (classes, properties, individuals) is accessible and resolvable on the web when clicking on the link.
Versioning of the schema
Each release of the SPHN RDF Schema has a version associated with it.
The version, indicated by the tag owl:versionIRI in the SPHN header information,
contains the year of release and the release number for that year
(e.g. https://biomedit.ch/rdf/sphn-schema/sphn/2024/2 corresponds to the second release
of the SPHN RDF Schema in 2024). The published Schema IRI will always point to the
latest version IRI of the SPHN RDF Schema.
The latest official release of the SPHN RDF Schema is: SPHN RDF Schema 2024.2
Previous versions can be found here.
SPHN header information
The header of the SPHN RDF Schema is contains the following information:
The title of the schema (
dc:title)A short description of the content of the schema (
dc:description)The license defining the terms and conditions of usage of the schema (
dcterms:license)The version of the schema (
owl:versionIRI)The external terminologies to be imported (
owl:imports)The previously released version of the schema (
owl:priorVersion)
Below is the header information of a turtle file:
<https://biomedit.ch/rdf/sphn-schema/sphn> a owl:Ontology ;
dc:description "The SPHN RDF Schema describing concepts defined in the official SPHN Dataset" ;
dc:rights "© Copyright 2024, Personalized Health Informatics Group (PHI), SIB Swiss Institute of Bioinformatics" ;
dc:title "The SPHN RDF Schema" ;
dcterms:bibliographicCitation "https://doi.org/10.1038/s41597-023-02028-y" ;
dcterms:license <https://creativecommons.org/licenses/by/4.0/> ;
owl:imports <http://purl.obolibrary.org/obo/eco/releases/2023-09-03/eco.owl>,
<http://purl.obolibrary.org/obo/genepio/releases/2023-08-19/genepio.owl>,
<http://purl.obolibrary.org/obo/geno/releases/2023-10-08/geno.owl>,
<http://purl.obolibrary.org/obo/obi/2023-09-20/obi.owl>,
<http://purl.obolibrary.org/obo/so/2021-11-22/so.owl>,
<http://snomed.info/sct/900000000000207008/version/20231201>,
<http://www.ebi.ac.uk/efo/releases/v3.61.0/efo.owl>,
<https://biomedit.ch/rdf/sphn-resource/atc/2024/1>,
<https://biomedit.ch/rdf/sphn-resource/chop/2024/4>,
sphn-edam:1.25,
<https://biomedit.ch/rdf/sphn-resource/emdn/2021-09-29/1>,
sphn-hgnc:20231215,
<https://biomedit.ch/rdf/sphn-resource/icd-10-gm/2024/3>,
<https://biomedit.ch/rdf/sphn-resource/loinc/2.76/1>,
<https://biomedit.ch/rdf/sphn-resource/ucum/2024/1>,
<https://www.orphadata.com/data/ontologies/ordo/last_version/ORDO_en_4.4.owl> ;
owl:priorVersion <https://biomedit.ch/rdf/sphn-ontology/sphn/2024/1> ;
owl:versionIRI <https://biomedit.ch/rdf/sphn-schema/sphn/2024/2> .
Note
The schema header provides information about the version of the external terminologies used in the SPHN RDF Schema 2024.2.
About the SPHN RDF classes
Naming convention for classes
The classes defined in the SPHN RDF Schema come from the concepts defined in the SPHN dataset
(column ‘general concept name’): one concept corresponds to one class. The unique identifier of
a class corresponds to a concatenation of the words forming the concept written in an UpperCase format.
For example, the concept Radiotherapy Procedure concept is defined as a RadiotherapyProcedure
class in RDF. A class necessarily contains the following information:
A
rdfs:labelcorresponding to the class name with a space between words for better readability,A
skos:definitioncontaining the description of the class.
A class may have other annotations which are detailed in the section Annotations for enriching knowledge content.
In addition, a few classes have a meaning binding associated with a
SNOMED CT, LOINC, GENO
or SO concept or code.
This meaning binding is represented in the schema with the annotation owl:equivalentClass.
Additional classes defined in the SPHN RDF Schema
Besides the concepts defined in the SPHN Dataset, four classes have been generated to represent specific types of metadata in RDF:
A
Terminologyclass: it groups classes of external resources made available in RDF (e.g. SNOMED CT, ATC, CHOP) and used within the SPHN context to be able to refer to them as possible values (i.e. the SNOMED CT code248152002is a possible value for the SPHNcodeproperty of theAdministrativeSex), or as equivalent classes (i.e. the LOINC code8867-4is an equivalent class of the SPHNHeartRateclass).A
ValueSetclass: it groups classes defining SPHN instances of possible values for certain object properties. The convention used for defining a ValueSet is:<Class>_<composedOf general concept name>(e.g.AdverseEvent_consequences,Sample_fixationType).
Note
The exception lies with the Comparator value set class which naming has been simplified in 2023.2
since the set of values used is the same in BirthDate and Quantity.
A
DataReleaseclass, which stamps the extraction date of a dataset together with information related to the version of the SPHN or project dataset (see versioning-of-the-data for more information).A
Deprecatedclass, which groups classes from a previous release of the schema that are no longer used in the current version.
Class hierarchies
Some hierarchies are defined in the SPHN Dataset in the column parent and interpreted in the RDF schema.
For instance, the class Diagnosis is a parent of BilledDiagnosis, ICDODiagnosis, NursingDiagnosis.
The rules of inheritance of properties is applied. All properties providing information about a parent class
are automatically inherited in the children class. Therefore, in the rdfs:Domain of
inherited properties, only the parent class is explicitly stated.
In general, all the properties are inherited from a parent class to a child class.
There is a distinction between the way the SPHN concepts and the project concepts are built. All SPHN
concepts, besides the deprecated ones, are subclasses to the class SPHNConcept (which is the root class of the SPHN RDF Schema).
Projects are able to extend SPHN concepts, meaning they can also create inherited concepts to SPHN concepts.
The projects have their own root class defined, PROJECTConcept.
To avoid any inheritance issue, this project’s root class (PROJECTConcept) and all external terminologies root classes
(which may be provided by the project) will be automatically added as children to the SPHNConcept.
When doing this, the root classes will be included in the
concept rdfs:Domain, which would result in inheritance of most properties in every single concept domain.
Hence, the root classes SPHNConcept and PROJECTConcept would be ignored for the inheritance by the SPHN tools, unless specified otherwise.
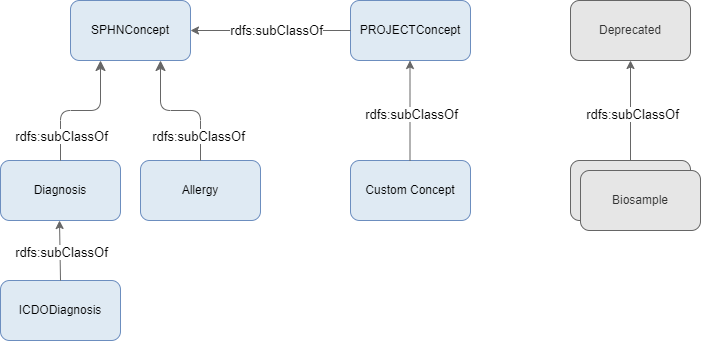
Figure 2 . SPHNConcept class (root of SPHN classes) is parent of a project’s root class.
About the SPHN RDF properties
Concepts in the SPHN Dataset contain compositions or so-called ‘composedOfs’ (i.e. information about a concept) that are translated in the SPHN RDF Schema as object properties (relationship between individuals) or data properties (relationship of individual to a literal data).
The IRIs of properties are named after the general concept name column in the SPHN Dataset.
The convention used is to define properties with has + <general concept name> as showcased in
the tables below (Table 2 for object properties and Table 3 for data properties).
Object properties
Object properties in the SPHN RDF Schema can define relationships between:
Resources from the clinical data (a given instance of
BodySiteis connected to a patient’sCircumferenceMeasurement) andResources and elements collected from external ontologies (the
Billed Diagnosisis identified by a specificICD-10-GMcode).
object property |
domain |
range |
|---|---|---|
hasBodySite |
CircumferenceMeasurement |
BodySite |
hasTherapeuticArea |
HealthcareEncounter |
TherapeuticArea |
hasCode |
BilledDiagnosis |
ICD-10-GM |
An additional set of five object properties are generated in the RDF schema, not represented in nor required by the SPHN Dataset:
hasSubjectPseudoIdentifier, connecting an information to the patient identifierhasDataProvider, connecting an information to the data providerhasAdministrativeCase, connecting an information to the administrative casehasSourceSystem, connecting an information to the source systemhasExtractionDateTime, providing information about the time of extraction of the data provided in a given RDF data file
Note
Since 2024, the same property can be reused in different context.
For instance, the property hasSample can be used to connect a data element to an instance of Sample
or to an instance of Isolate, which is in this case a descendant of Sample in SPHN.
Another example is hasResult which can connect to any instance of a Result.
In the case of a Blood Pressure Measurement, for instance, hasResult points to an instance of Blood Pressure only.
The applicable range of a property for a given context is encoded and specified
in the SPHN RDF Schema with the restrictions.
Data properties
Data properties in the SPHN RDF Schema point to literal values of given SPHN concepts.
data property |
general concept name (from SPHN Dataset) |
domain |
range |
|---|---|---|---|
hasValue |
value |
Quantity |
xsd:double |
hasStandardGuideline |
standard guideline |
Interpretation |
xsd:string |
hasRecordDateTime |
record datetime |
Excluded Disorder |
xsd:datetime |
Constraints added to properties
An owl:Restriction is a particular type of class description that puts value
and cardinality constraints to a property.
There can be multiple constraints added to narrow down and/or enrich a concept.
Value constraints are meant to restrict the possible types of values of a property.
Cardinality constraints restrict the number of instances of a property.
The rules for inheritance of restrictions are applied. Any restrictions annotated on a property are automatically inherited by the subproperties. That is, if a subclass (inheriting the parent class’ property) has a subproperty of that property, the restrictions of this subproperty must be in the range of the restrictions of the parent property. For example, if the parent property has cardinality 0:1, the subproperty may have cardinalities 0:1 or 1:1.
Each instance of a subproperty is also an instance of the parent property.
Value constraint
The range of a property usually encodes the type of value allowed for this property. However, in some cases, it was not possible to straightforwardly encode additional information needed to account for the full context. In these cases, value constraints are used to ensure a clean modeling of the data.
For instance, the Code of a Unit given to an OxygenSaturation is %.
However, it is not possible to annotate that the property hasCode can only have as a
range % since hasCode can be used in another context.
To solve this issue, value constraints (using owl:Restriction) are used.
Coming back to the example of the oxygen saturation’s unit constraint, the following constraint (in .ttl format) can be built:
sphn:OxygenSaturation rdf:type owl:Class ;
owl:subClassOf [ rdf:type owl:Restriction ;
owl:onProperty sphn:hasQuantity
owl:someValuesFrom [ rdf:type owl:Restriction
owl:onProperty sphn:hasUnit ;
owl:someValuesFrom [ rdf:type owl:Restriction ;
owl:onProperty sphn:hasCode ;
owl:someValuesFrom ucum:percent ]]];
The code above can be read line by line as:
The
sphn:OxygenSaturationis a class.It has the following restriction statement:
whenever the
sphn:hasQuantityis used for describing thesphn:OxygenSaturation,only some specific values are allowed, which is in reality a second constraint statement.
A constraint is applied to the property
sphn:hasUnitthat again also has only specific values allowed leading to a third constraint statement.
A constraint is now applied to the property
sphn:hasCodewhich allows only values that are:
ucum:percent.
In other words, when annotating the quantity of an OxygenSaturation,
the Code representing the Unit must be the instance of percent
coming from the UCUM notation (see Figure 3). No other values are allowed.
The constraints are represented in a nested structure that follows the properties’
path as they are encoded in the SPHN RDF Schema.
Figure 3 . Example of property path from OxygenSaturation to the possible code of the Unit.
Cardinality constraint
Cardinalities constraints have been implemented to restrict the number of values an instance of
a class may have for specific properties. The owl:minCardinality and owl:maxCardinality
notation have been used.
For instance, the Sample must at least one (minCardinality) and maximum one (maxCardinality) collection datetime connected:
owl:intersectionOf ( [ a owl:Restriction ;
owl:minCardinality "1"^^xsd:nonNegativeInteger ;
owl:onProperty sphn:hasCollectionDateTime ] [ a owl:Restriction ;
owl:maxCardinality "1"^^xsd:nonNegativeInteger ;
owl:onProperty sphn:hasCollectionDateTime ] ) ],
Both value and cardinality constraints are visualized in the PyLODE documentation in the definitions of the classes (https://biomedit.ch/rdf/sphn-schema/sphn/2024/2).
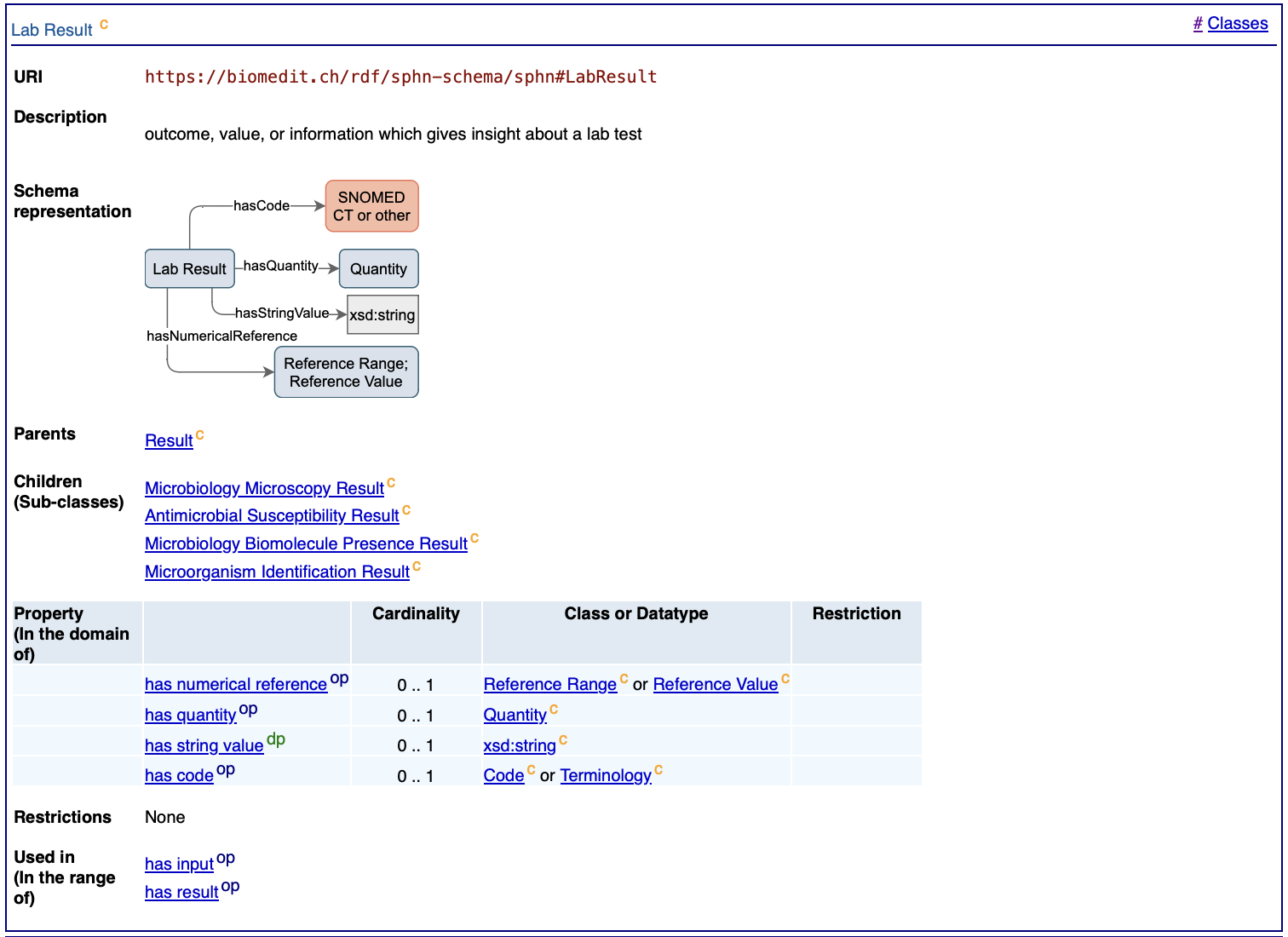
Figure 4. Example of cardinality constraints shown in the SPHN RDF visualization for Lab Result (with PyLODE).
Annotations for enriching knowledge content
In some classes, there is a need to annotate specific information provided in the SPHN Dataset. These additional annotations relate to exceptions in the way property restrictions should be understood. Therefore, a new SPHN annotation was created to ease connecting elements from one SPHN RDF Schema to another.
skos:scopeNote for subclass values not allowed
SNOMED CT provides a hierarchical structure of its codes.
When values are given as SNOMED CT codes, it implicitly means that,
by default, any children of the provided code are valid results when taking
into account the inheritance rules defined in Semantic Web: a child class
is a parent class. The child class is simply a more specific element of the parent class.
Sometimes SPHN makes restrictions and only allows for explicitly stated codes
to be valid in a defined value set.
This information is encoded as a skos:scopeNote with the following text
provided for each property: sphn:<property> no subclasses allowed.
For instance, values for defining the administrative gender of a patient are strictly limited
to male, female, indeterminate. This value set is provided as an owl:Restriction:
[ rdf:type owl:Restriction ;
owl:onProperty sphn:hasCode ;
owl:someValuesFrom [ rdf:type owl:Class ;
owl:unionOf ( snomed:248152002
snomed:248153007
snomed:32570681000036106
)
]
]
In addition, the following annotation, given as information in the schema, states that only these values are permitted and no children of these codes are allowed:
sphn:AdministrativeSex skos:scopeNote "sphn:hasCode no subclasses allowed" .
This information is used in the SHACLer for strictly limiting the set of values of a
property and is provided in the PyLODE visualization of the SPHN RDF Schema to indicate
in the restriction field that child terms are not allowed. Here is the same AdministrativeSex
hasCode property example represented in PyLODE:

Figure 5. Example of the interpretation of a skos:scopeNote constraint in the SPHN RDF visualization (with PyLODE). The skos:scopeNote indicates that the child terms of the given value set are not allowed to be provided.
skos:note for permitted standards and extendable value sets
The SPHN dataset in some cases defines the external standards (or terminologies) that can be used
for a given property in a certain context. This information is encoded in the schema as a skos:note.
For instance, the code of a ProblemCondition may come from ICPC or another standard.
This information is depicted as follow in .ttl:
sphn:ChromosomalLocation skos:note "sphn:hasEndCytobandCode allowed coding system: ISCN" .
The SPHN dataset in some cases also defines extendable value sets.
Extendable value sets establish properties with recommended or example values,
where the data provider has the option of extending them with other values as they see fit.
This knowledge is represented as follows in RDF: the owl:Restriction given for this
property value is not taking into account the single values provided by the dataset but
matches to a higher level of class in the hierarchy from the coding system, enabling other values to be used.
The values recommended in the SPHN dataset are provided with the annotation skos:note.
For instance, the list of codes about body sites where the oxygen saturation can be measured
is an extendable value set in SPHN. The owl:Restriction is given in BodySite where
Code values allowed are only children of snomed:123037004 |body structure (body structure)|,
as represented below in .ttl:
[ rdf:type owl:Restriction ;
owl:onProperty sphn:hasCode ;
owl:someValuesFrom <http://snomed.info/id/123037004>
]
The specific values recommended in the SPHN dataset to be used are:
snomed:29707007 |Toe structure (body structure)|, snomed:7569003 |Finger structure (body structure)|,
snomed:48800003 |Ear lobule structure (body structure)|.
These values are indicated in the following note annotated in the OxygenSaturation class:
sphn:OxygenSaturation skos:note "sphn:hasBodySite/sphn:hasCode recommended values [snomed:29707007; snomed:7569003; snomed:48800003]"
sphn:replaces for reference of previous classes and properties
Changes in classes or properties between different versions of the
SPHN RDF Schema are tracked directly in RDF with the sphn:replaces annotation.
For instance, in SPHN 2022.1, the property https://biomedit.ch/rdf/sphn-schema/sphn#hasActiveIngredient
replaced the property https://biomedit.ch/rdf/sphn-schema/sphn#hasDrugActiveIngredientSubstance
defined in SPHN 2021.2. The following statement is written in the SPHN 2022.1 RDF schema:
sphn:hasActiveIngredient rdf:type owl:ObjectProperty ;
sphn:replaces sphn:hasDrugActiveIngredientSubstance .
This annotation is currently used in the migration path tool to generate a CSV file of the differences between two versions of SPHN RDF Schema.
Development process
After the release of each new version of the SPHN Dataset the corresponding SPHN RDF Schema is reviewed by the SPHN Data Coordination Center (DCC) in close collaboration with the IT experts of the Swiss University Hospitals and revised as necessary.
Availability and usage rights
The SPHN RDF Schema is available at https://git.dcc.sib.swiss/sphn-semantic-framework/sphn-schema/, a HTML-based decription of the concepts can be found https://biomedit.ch/rdf/sphn-schema/sphn as well as a interactivly browseable Schema on https://schemascope.dcc.sib.swiss/.
External terminologies are accessible through the Terminology Service.
If you need further information, please contact the SPHN Data Coordination Center (DCC) at dcc@sib.swiss.
The SPHN RDF Schema is under the CC BY 4.0 License.
Further reading
Touré, V., Krauss, P., Gnodtke, K., Buchhorn, J., Unni, D., Horki, P., Raisaro, J.L., Kalt, K., Teixeira, D., Crameri, K. and Österle, S. (2023). FAIRification of health-related data using semantic web technologies in the Swiss Personalized Health Network. Scientific Data, 10(1), p.127 (doi: 10.1038/s41597-023-02028-y)